SimPhoNy¶
Mayavi tools are available in the simphony library through the
visualisation plug-in named mayavi_tools.
e.g:
from simphony.visualisation import mayavi_tools
Visualizing CUDS¶
The show() function is available to
visualise any top level CUDS container. The function will open a
window containing a 3D view and a mayavi toolbar. Interaction
allows the common mayavi operations.
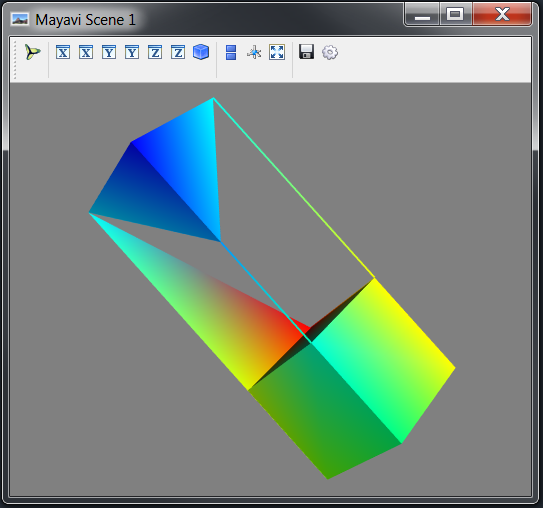
Mesh example
from numpy import array
from simphony.cuds.mesh import Mesh, Point, Cell, Edge, Face
from simphony.core.data_container import DataContainer
points = array([
[0, 0, 0], [1, 0, 0], [0, 1, 0], [0, 0, 1],
[2, 0, 0], [3, 0, 0], [3, 1, 0], [2, 1, 0],
[2, 0, 1], [3, 0, 1], [3, 1, 1], [2, 1, 1]],
'f')
cells = [
[0, 1, 2, 3], # tetra
[4, 5, 6, 7, 8, 9, 10, 11]] # hex
faces = [[2, 7, 11]]
edges = [[1, 4], [3, 8]]
mesh = Mesh('example')
# add points
uids = [
mesh.add_point(
Point(coordinates=point, data=DataContainer(TEMPERATURE=index)))
for index, point in enumerate(points)]
# add edges
edge_uids = [
mesh.add_edge(
Edge(points=[uids[index] for index in element]))
for index, element in enumerate(edges)]
# add faces
face_uids = [
mesh.add_face(
Face(points=[uids[index] for index in element]))
for index, element in enumerate(faces)]
# add cells
cell_uids = [
mesh.add_cell(
Cell(points=[uids[index] for index in element]))
for index, element in enumerate(cells)]
if __name__ == '__main__':
from simphony.visualisation import mayavi_tools
# Visualise the Mesh object
mayavi_tools.show(mesh)
Lattice example
import numpy
from simphony.cuds.lattice import make_cubic_lattice
from simphony.core.cuba import CUBA
lattice = make_cubic_lattice('test', 0.1, (5, 10, 12))
for node in lattice.iter_nodes():
index = numpy.array(node.index) + 1.0
node.data[CUBA.TEMPERATURE] = numpy.prod(index)
lattice.update_node(node)
if __name__ == '__main__':
from simphony.visualisation import mayavi_tools
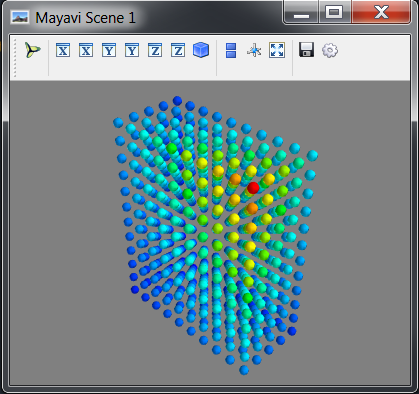
# Visualise the Lattice object
mayavi_tools.show(lattice)
Particles example
from numpy import array
from simphony.cuds.particles import Particles, Particle, Bond
from simphony.core.data_container import DataContainer
points = array([[0, 0, 0], [1, 0, 0], [0, 1, 0], [0, 0, 1]], 'f')
bonds = array([[0, 1], [0, 3], [1, 3, 2]])
temperature = array([10., 20., 30., 40.])
particles = Particles('test')
uids = []
for index, point in enumerate(points):
uid = particles.add_particle(
Particle(
coordinates=point,
data=DataContainer(TEMPERATURE=temperature[index])))
uids.append(uid)
for indices in bonds:
particles.add_bond(Bond(particles=[uids[index] for index in indices]))
if __name__ == '__main__':
from simphony.visualisation import mayavi_tools
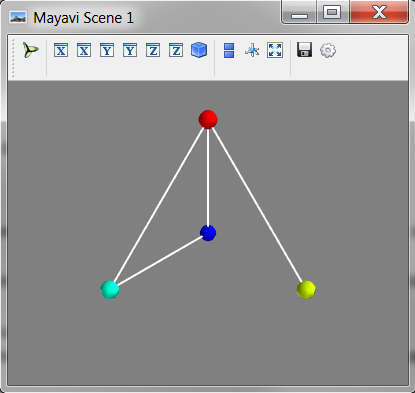
# Visualise the Particles object
mayavi_tools.show(particles)
Create VTK backed CUDS¶
Three objects (i.e class:~. VTKMesh, class:~.VTKLattice, ~.VTKParticles) that wrap a VTK dataset and provide the CUDS top level container API are also available. The vtk backed objects are expected to provide memory and some speed advantages when Mayavi aided visualisation and processing is a major part of the working session. The provided examples are equivalent to the ones in section Visualizing CUDS.
Note
Note all CUBA keys are supported for the data attribute of the contained items. Please see documentation for more details.
VTK Mesh example
from numpy import array
from simphony.cuds.mesh import Point, Cell, Edge, Face
from simphony.core.data_container import DataContainer
from simphony.visualisation import mayavi_tools
points = array([
[0, 0, 0], [1, 0, 0], [0, 1, 0], [0, 0, 1],
[2, 0, 0], [3, 0, 0], [3, 1, 0], [2, 1, 0],
[2, 0, 1], [3, 0, 1], [3, 1, 1], [2, 1, 1]],
'f')
cells = [
[0, 1, 2, 3], # tetra
[4, 5, 6, 7, 8, 9, 10, 11]] # hex
faces = [[2, 7, 11]]
edges = [[1, 4], [3, 8]]
mesh = mayavi_tools.VTKMesh('example')
# add points
uids = [
mesh.add_point(
Point(coordinates=point, data=DataContainer(TEMPERATURE=index)))
for index, point in enumerate(points)]
# add edges
edge_uids = [
mesh.add_edge(
Edge(points=[uids[index] for index in element]))
for index, element in enumerate(edges)]
# add faces
face_uids = [
mesh.add_face(
Face(points=[uids[index] for index in element]))
for index, element in enumerate(faces)]
# add cells
cell_uids = [
mesh.add_cell(
Cell(points=[uids[index] for index in element]))
for index, element in enumerate(cells)]
if __name__ == '__main__':
# Visualise the Mesh object
mayavi_tools.show(mesh)
VTK Lattice example
import numpy
from simphony.core.cuba import CUBA
from simphony.visualisation import mayavi_tools
lattice = mayavi_tools.VTKLattice.empty(
'test', 'Cubic', (0.1, 0.1, 0.1), (5, 10, 12), (0, 0, 0))
for node in lattice.iter_nodes():
index = numpy.array(node.index) + 1.0
node.data[CUBA.TEMPERATURE] = numpy.prod(index)
lattice.update_node(node)
if __name__ == '__main__':
# Visualise the Lattice object
mayavi_tools.show(lattice)
VTK Particles example
from numpy import array
from simphony.core.data_container import DataContainer
from simphony.cuds.particles import Particle, Bond
from simphony.visualisation import mayavi_tools
points = array([[0, 0, 0], [1, 0, 0], [0, 1, 0], [0, 0, 1]], 'f')
bonds = array([[0, 1], [0, 3], [1, 3, 2]])
temperature = array([10., 20., 30., 40.])
particles = mayavi_tools.VTKParticles('test')
uids = []
for index, point in enumerate(points):
uid = particles.add_particle(
Particle(
coordinates=point,
data=DataContainer(TEMPERATURE=temperature[index])))
uids.append(uid)
for indices in bonds:
particles.add_bond(Bond(particles=[uids[index] for index in indices]))
if __name__ == '__main__':
# Visualise the Particles object
mayavi_tools.show(particles)
Adapting VTK datasets¶
The adapt2cuds() function is
available to wrap common VTK datsets into top level CUDS
containers. The function will attempt to automatically adapt the
(t)vtk Dataset into a CUDS container. When automatic conversion
fails the user can always force the kind of the container to adapt into.
Furthermore, the user can define the mapping of the included
attribute data into corresponding CUBA keys (a common case for
vtk datasets that come from vtk reader objects).
Example
from numpy import array, random
from tvtk.api import tvtk
from simphony.core.cuba import CUBA
from simphony.visualisation import mayavi_tools
def create_unstructured_grid(array_name='scalars'):
points = array(
[[0, 1.2, 0.6], [1, 0, 0], [0, 1, 0], [1, 1, 1], # tetra
[1, 0, -0.5], [2, 0, 0], [2, 1.5, 0], [0, 1, 0],
[1, 0, 0], [1.5, -0.2, 1], [1.6, 1, 1.5], [1, 1, 1]], 'f') # Hex
cells = array(
[4, 0, 1, 2, 3, # tetra
8, 4, 5, 6, 7, 8, 9, 10, 11]) # hex
offset = array([0, 5])
tetra_type = tvtk.Tetra().cell_type # VTK_TETRA == 10
hex_type = tvtk.Hexahedron().cell_type # VTK_HEXAHEDRON == 12
cell_types = array([tetra_type, hex_type])
cell_array = tvtk.CellArray()
cell_array.set_cells(2, cells)
ug = tvtk.UnstructuredGrid(points=points)
ug.set_cells(cell_types, offset, cell_array)
scalars = random.random(points.shape[0])
ug.point_data.scalars = scalars
ug.point_data.scalars.name = array_name
scalars = random.random((2, 1))
ug.cell_data.scalars = scalars
ug.cell_data.scalars.name = array_name
return ug
# Create an example
vtk_dataset = create_unstructured_grid()
# Adapt to a mesh by converting the scalars attribute to TEMPERATURE
container = mayavi_tools.adapt2cuds(
vtk_dataset, 'test',
rename_arrays={'scalars': CUBA.TEMPERATURE})
if __name__ == '__main__':
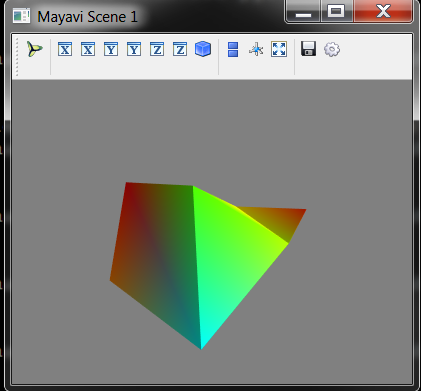
# Visualise the Lattice object
mayavi_tools.show(container)
Loading into CUDS¶
The load() function is available to
load mayavi readable files (e.g. VTK xml format) into top level
CUDS containers. Using load the user can import inside their
simulation scripts files that have been created by other
simulation application and export data into one of the Mayavi
supported formats.
